Help! How do I do a plot in R? (Part 2)
In this post we are going to see the different types of graphics that we can make with base R depending on the information we need to convey.

Welcome to the fourth installment of this saga titled How to analyze quantitative data without dying trying. In previous installments we saw how to start viewing our data in RStudio, how to calculate basic measurements of descriptive statistics and how to generate automatic graphs. In this post we are going to go a little deeper into the types of graphs that we can make, no longer automatically, but depending on the type of information we want to transmit.
Data visualization is a fairly important topic today both in the world of data science and in scientific communication. Although it seems that there are as many opinions as there are people who generate them, my position is that a good graph is not always the most striking or complex, but rather the one that best suits the communicative context and the one that makes our lives the most easier. There are social spheres where graphics are valued for their ability to present various dimensions of a phenomenon in a small space, such as in the world of infographics, but there are others where the graphic is a complement to other types of information, such as a scientific article. This context will define how much information we will have to condense in the graph and how much time we can dedicate to it.
There are many R packages that provide very powerful and complex tools for generating graphs, such as the undisputed king {ggplot2} or {plotly}. However, when we start learning to program in R, these complex tools become lifesavers, not only because of the wide variety of options, but also because the learning curve becomes a bit steep. This is why here we are going to concentrate on the graphics that we can make with base R and always focusing on what we want to convey, rather than on the graph itself.
First step: how to think about a graph?
As in previous posts, we are going to work with penguin data:
penguins <- palmerpenguins::penguinsIn the previous post we saw an intuitive way to start graphing the data, thanks to the plot() function that evaluates the variables we choose and based on that decides what type of graph to make. This comes in handy when we are starting out and are in an exploratory stage of the data. However, in real research, we are going to start defining questions that we want the data to answer us; Sometimes these questions are prior to the data and have to do with the research design and sometimes these questions arise from the exploration itself.
In general, it is advisable to start with a research question and then think about what graph can provide us with that answer. However, in order to reach that stage we have to know in advance what types of graphs exist in order to realize which one we need. For that reason alone, this post is going to be organized based on the types of graphs, instead of focusing on the questions.
The vast majority of graphs have two axes, the X (horizontal) and the Y (vertical); This means that in principle we can think of the graph as showing us how two variables or two aspects of our data are related. That is the starting point, always: we want to see something (an axis) in terms of another something (another axis). By convention, the X axis is used for the independent variable (that is, what is given) and the Y axis for the dependent or outcome variable (that it, what will change in terms of that independent variable). Then, on that basis we will be able to make the information more complex and specify other crossings.
Finally, another aspect to keep in mind is that the type of variable we have will restrict the different types of graphs we can choose. This means that numerical and categorical variables, for example, will give rise to different graphs (you can explore a little more here). I know, I know that this seems like a lot of information to you, but I assure you that many of these questions will become intuitive over time. Let us begin!
The histogram
The histogram is one of the least common graphs in academic articles but at the same time most important in practically everything we do in statistics. Basically it is a graph that shows how a variable is distributed in the data set that makes up our sample. Take a single numerical variable and specify how many elements in the sample have a particular value. For example:
hist(penguins$bill_length_mm)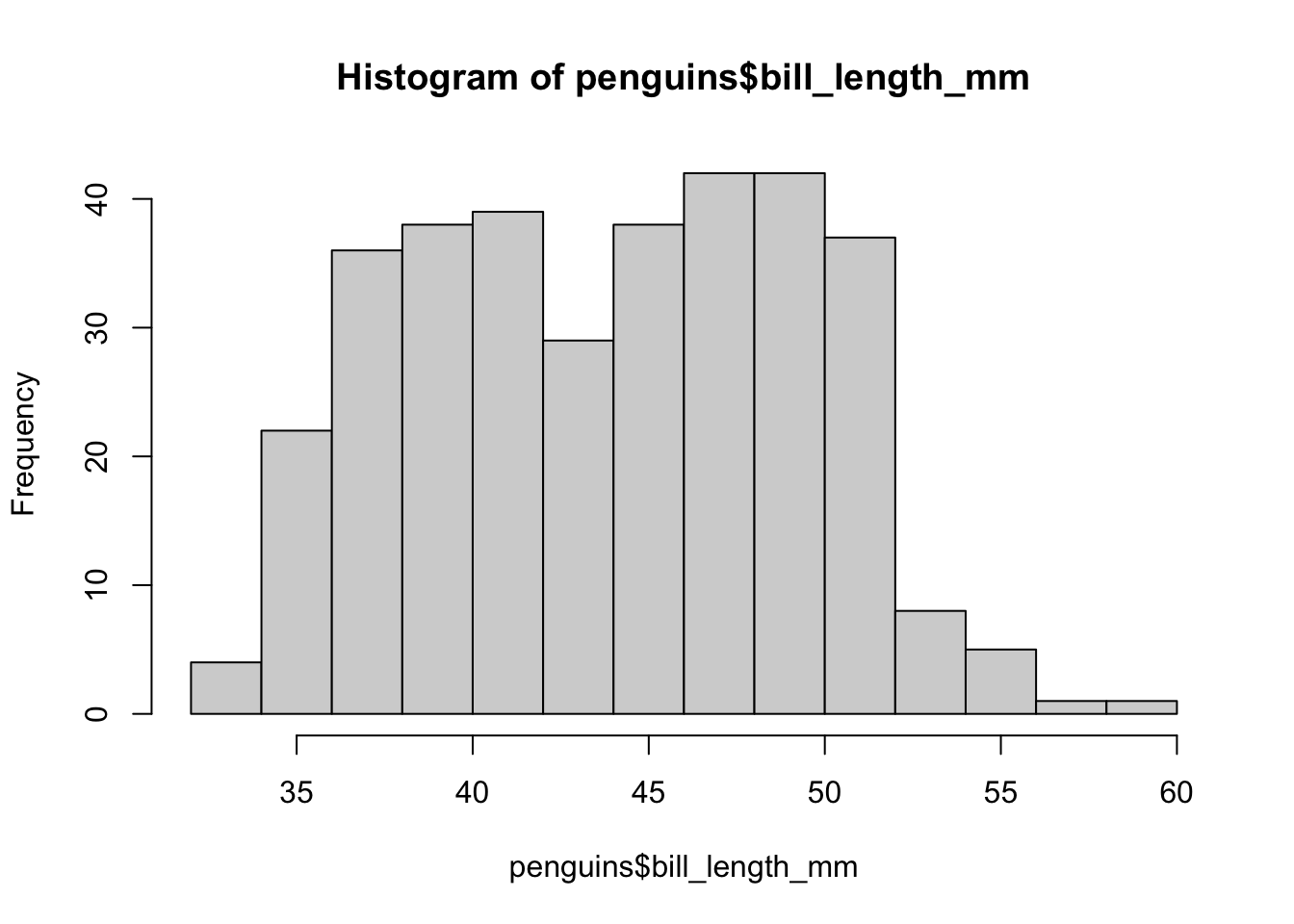
This graph shows us on the X axis the values that the length of the beak of the penguins in the sample can take, and in the Y axis the amount of penguins whose beaks have that length. We then see that there seem to be two large groups: the first with peaks between 30 and 43 mm, and the second between 43 and 60mm. This graph allows us to see which peak length values are more frequent (around 40 and around 50) and which are rare (less than 35 and more than 55).
If we think about it visually, we see that a histogram is a set of bars that mark quantities. The width of each bar represents an interval within the possible lengths, but in this case it does not show a precise value. In fact, we can manually modify the number of bars that the histogram will have with the breaks argument:
hist(penguins$bill_length_mm, breaks = 5)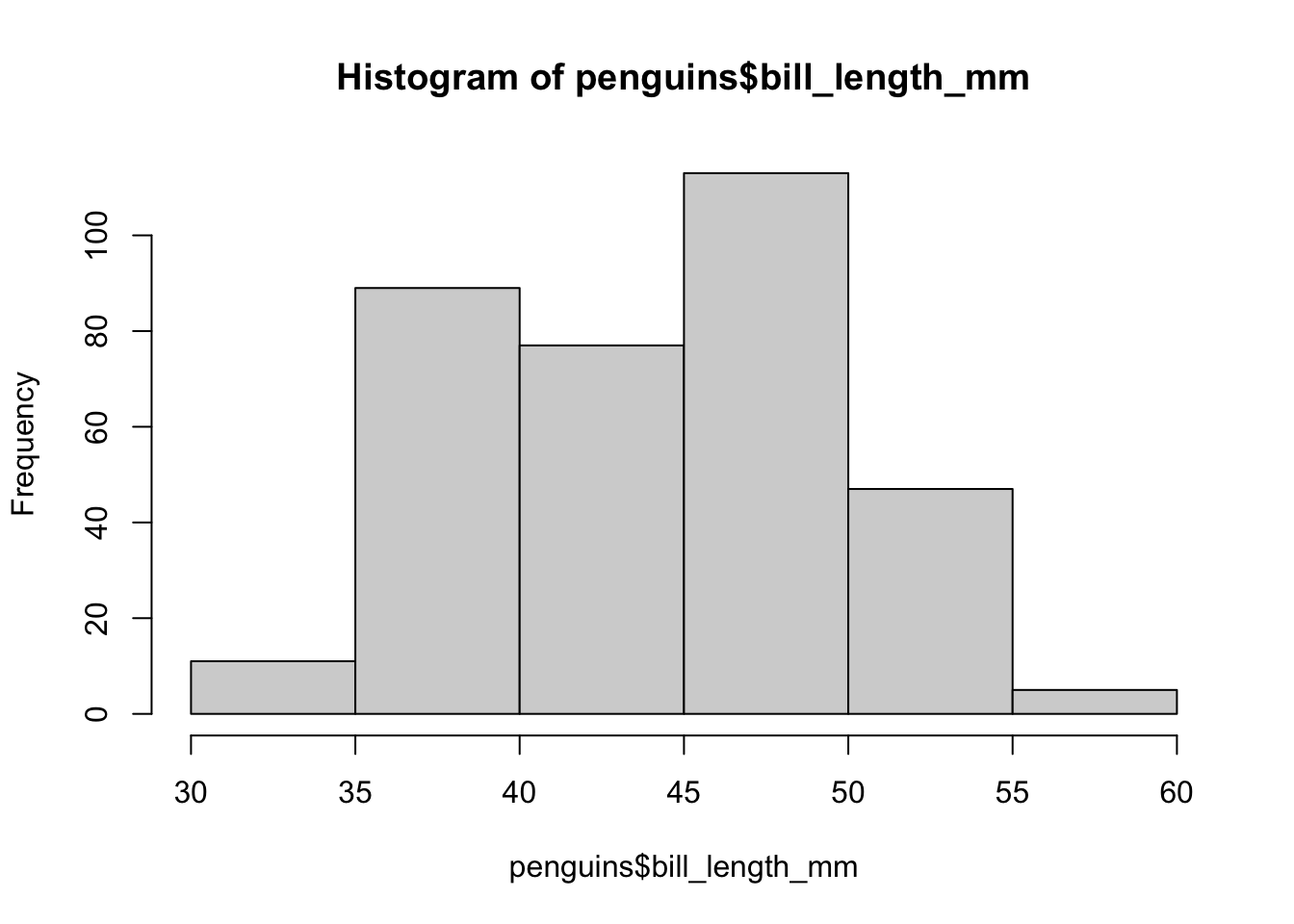
In this case we reduced the number of bars compared to the previous version and the result is very different: although we see in more detail that 35mm is an important breaking value and that most penguins have beaks between 45 and 50mm long, we lost precision and sensitivity with respect to the groups. In fact, in the previous post we saw that the link between the length and height of the beak showed three differentiated groups, so it would be logical to expect to see something like this in this graph. We can try other bar values:
hist(penguins$bill_length_mm, breaks = 50)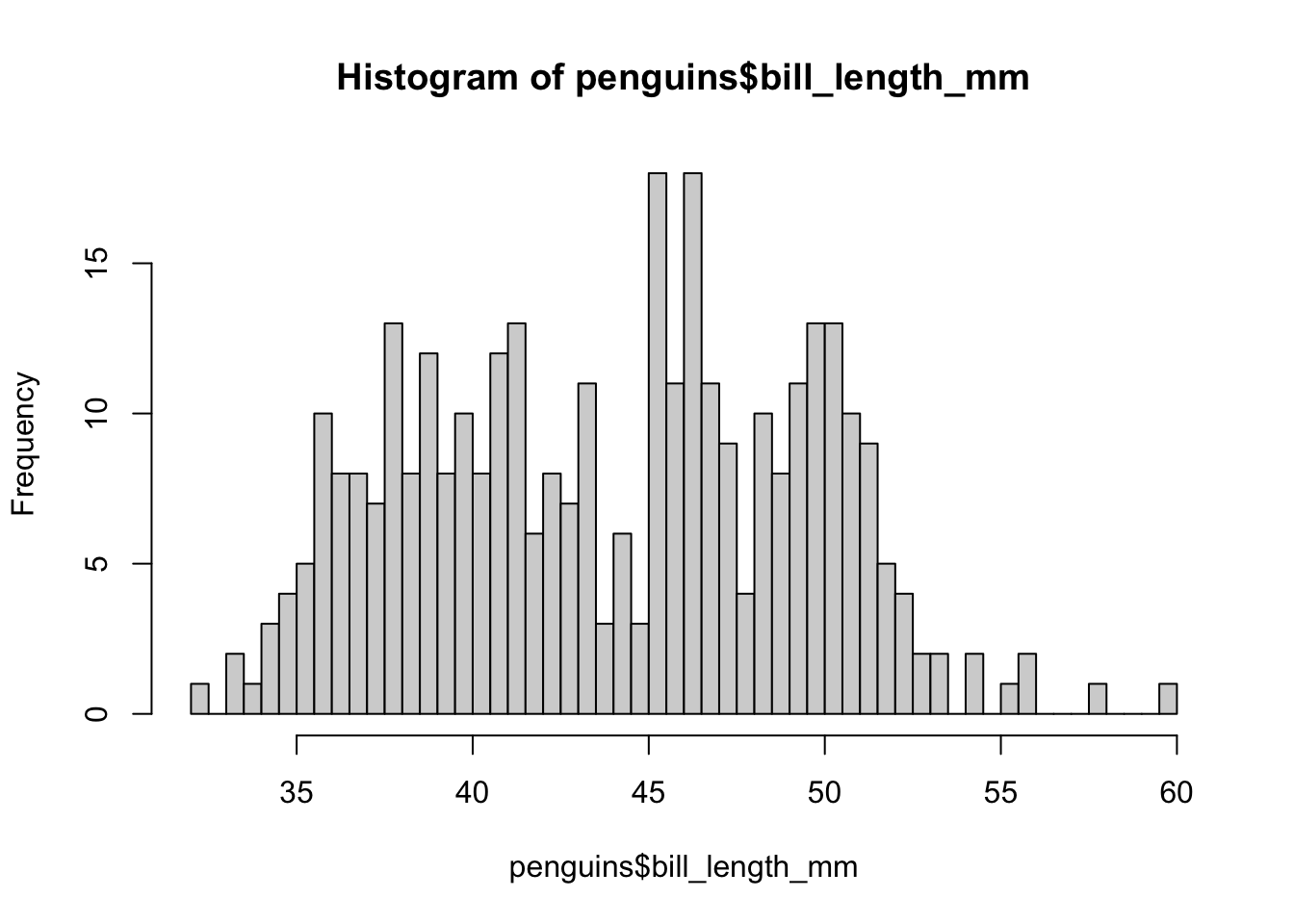
hist(penguins$bill_length_mm, breaks = 100)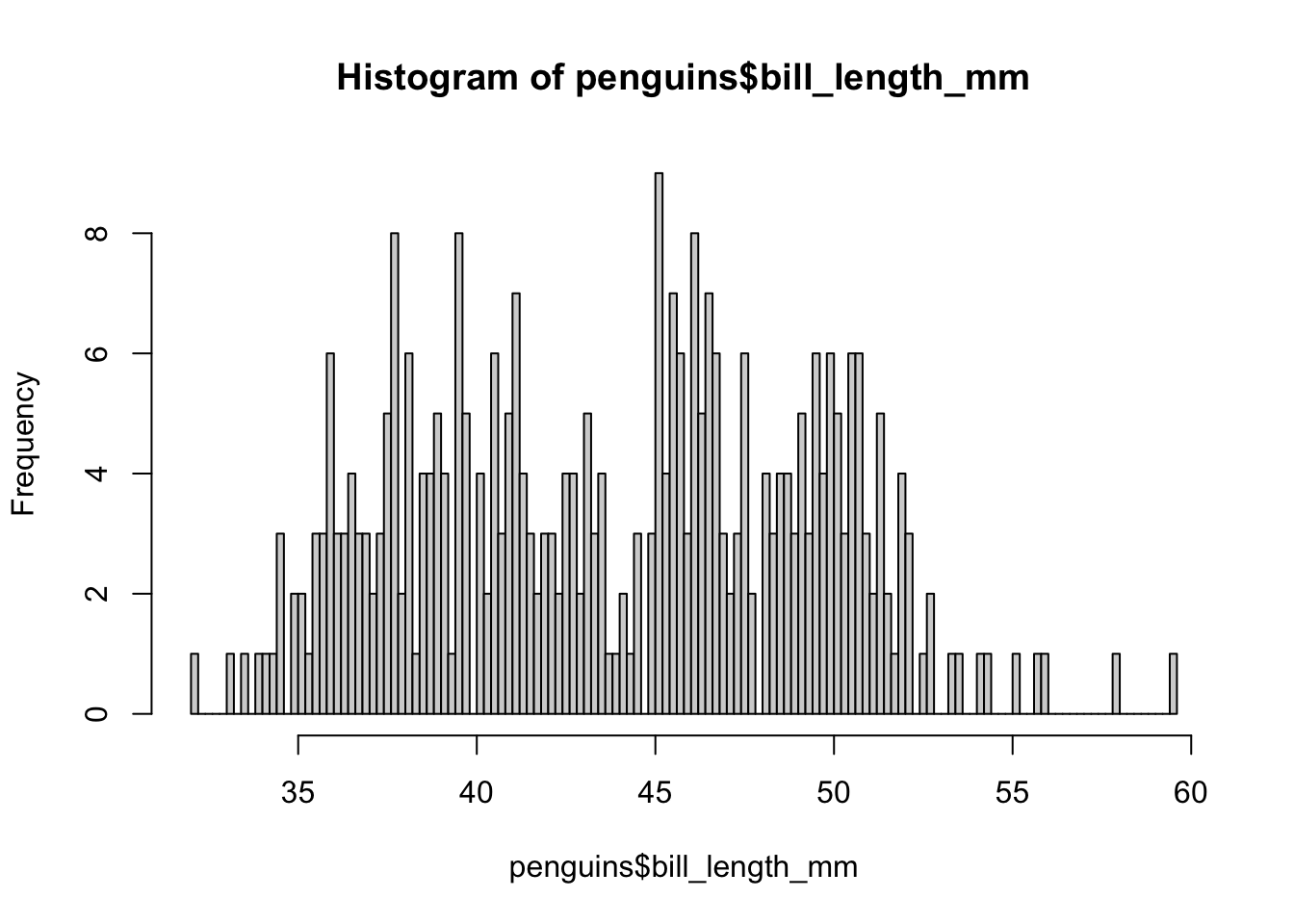
hist(penguins$bill_length_mm, breaks = 200)
As we increase the number of bars, we approach one bar per penguin, which is perhaps not the most interesting or necessary, because already at 50 we begin to notice the differentiation between groups. Sometimes it is a matter of testing what number of bars is the most appropriate for what we want to graph.
We then find that there are three large groups according to which the length of the beak varies. We look at our data and see that there are three species of penguins: Adelie, Chinstrap, and Gentoo, so we wonder if the length of the beaks could be due to that. Luckily, we have a graph that can help us: the bar graph.
The bar chart
This has to be the most common chart in the world of data visualization. Some people look down on it for its simplicity, but the truth is that it can be very useful and easy to read (although it has some biases, as we will see later). So, I want to know if the length of the beaks varies depending on the species of penguins. For that, I first need to calculate the averages for each of the species. You will remember that in the previous post we used the aggregate function; the problem is that the result is an object that we cannot easily use to graph. We can use the tapply function which basically does the following: on the object X, apply the FUN function from the groups generated by INDEX1. And to that table, we apply the barplot() function to generate the bar graph:
barplot(tapply(X = penguins$bill_length_mm,
INDEX = penguins$species,
FUN = mean, na.rm = TRUE))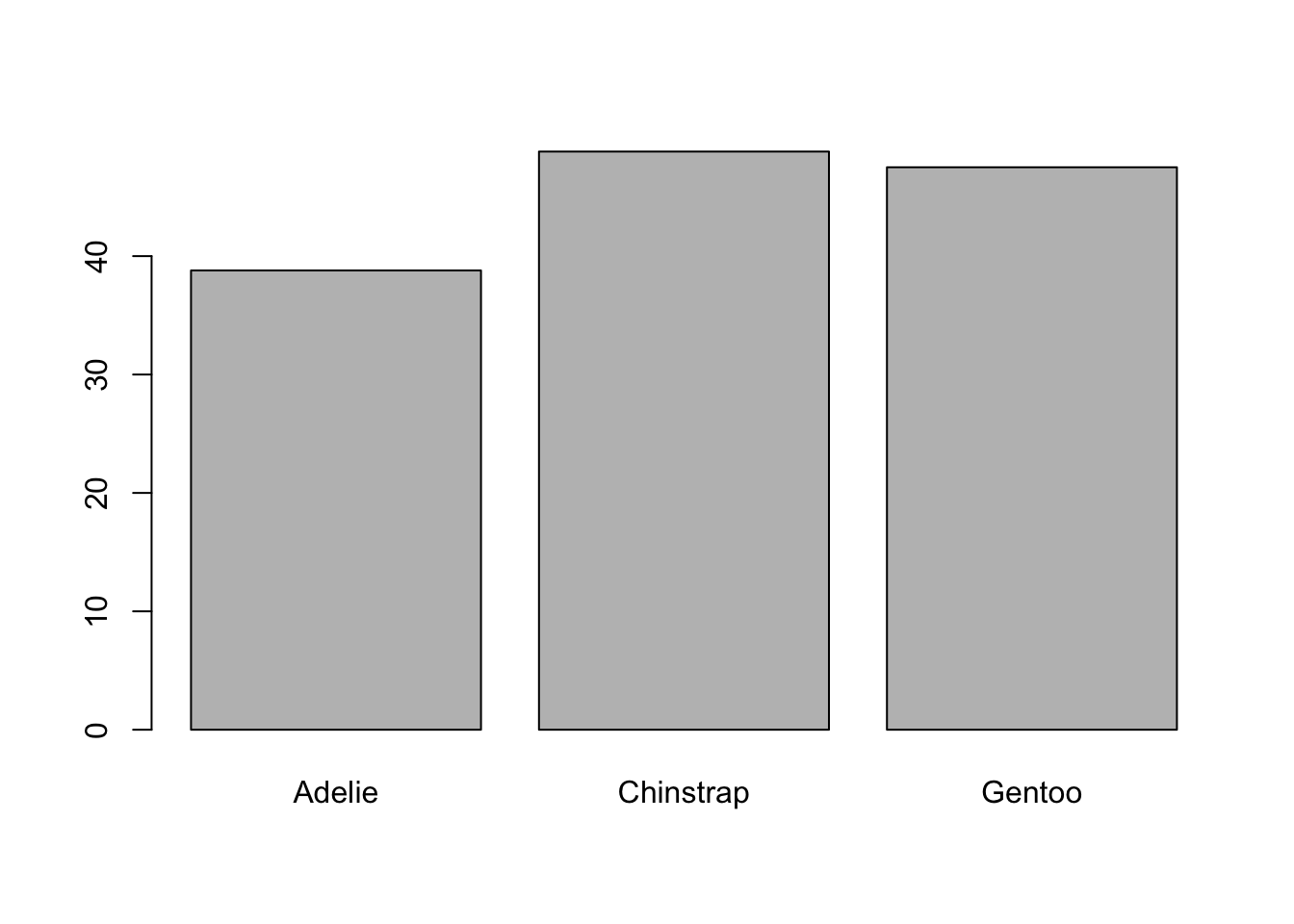
Programming languages allow this type of recursion (one thing inside another) very easily, as long as the object that generates the first function (tapply()) is an object that the second function (barplot() ) can take. Anyway, for us flesh and blood people something like that can become a little difficult to read; An alternative is to split this into two steps, first creating an object with the means and then plotting the object:
# first we create the table
mean_datatable <- tapply(X = penguins$bill_length_mm,
INDEX = penguins$species,
FUN = mean, na.rm = TRUE)
# then we graph the table
barplot(mean_datatable)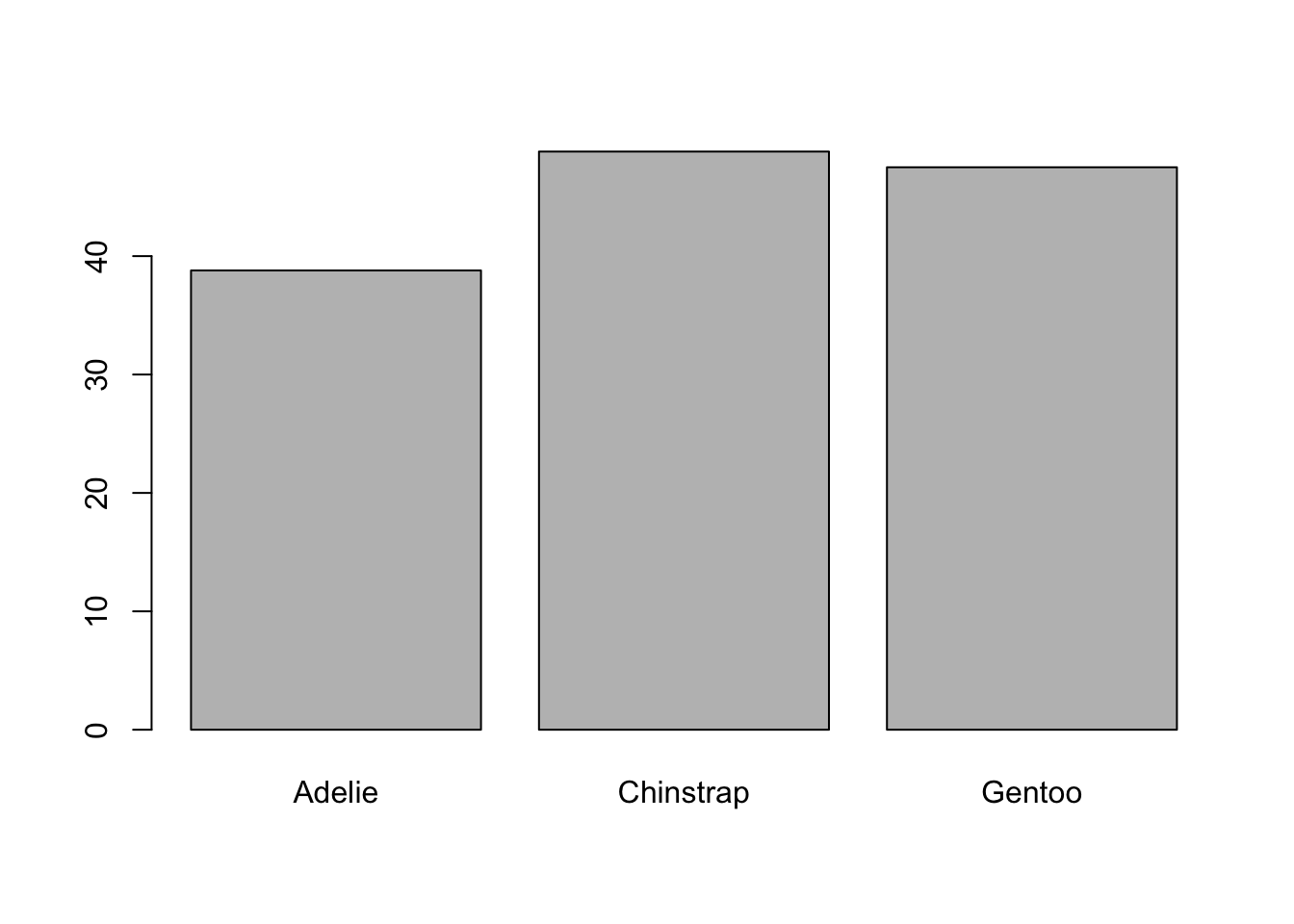
The result is the same. In general, I am not a big fan of creating many intermediate objects that I will not use again later, because I consider that it leads to confusion, but it is true that it helps make the code easy to read.
Let’s go back to the graph. As we saw in the previous post we can add a title to the graph or modify the axis labels. Beyond that, there is something that should have caught your attention: if you look at the Y-axis ruler, you will see that it cuts at 40. Why? I don’t know, sometimes it happens. We can correct this by specifying an argument called ylim, which defines the numerical limits between which the graph extends. Since there are two values, we must concatenate them with the c() function:
barplot(tapply(X = penguins$bill_length_mm,
INDEX = penguins$species,
FUN = mean, na.rm = TRUE),
ylim = c(0, 55))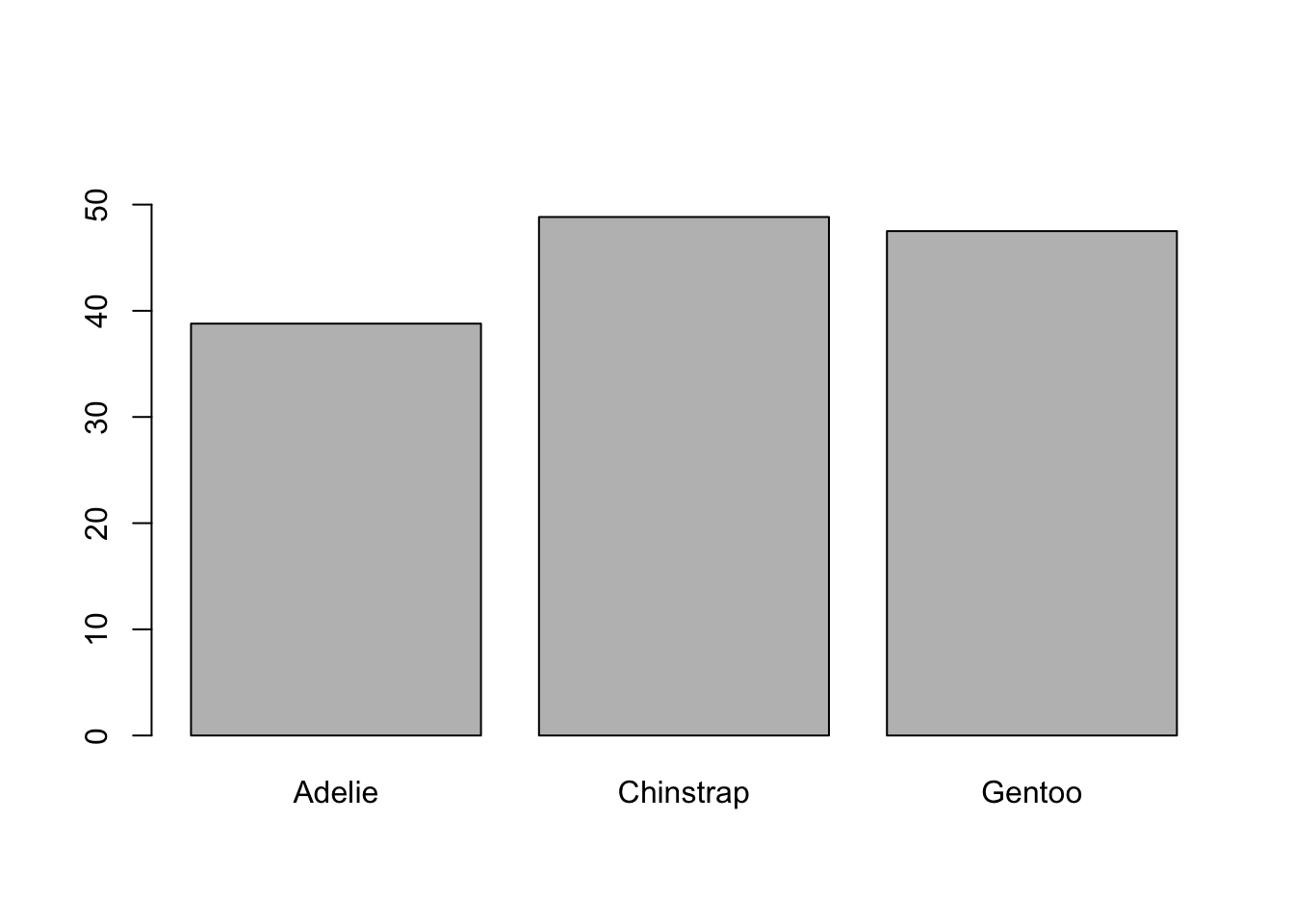
Well, we see then that there are some differences between the beak length of penguins depending on the species, especially between Adelia and Chinstrap. However, graphing means in this way has a bias that we talked about previously: the mean is a measure that is very sensitive to extreme values and at the same time hides them, because it does not allow us to see the existing variation (which we know exists, because we saw it in the histogram). For that we have another graph: the boxplot.
The boxplot
The boxplot is a type of graph that directly represents the following statistical measures: median, quartiles, interquartile range and outliers. You can read a little more about this type of graph on its Wikipedia site, but the most important thing is that it allows us to see the dispersion of the data. Let’s see:
boxplot(penguins$bill_length_mm)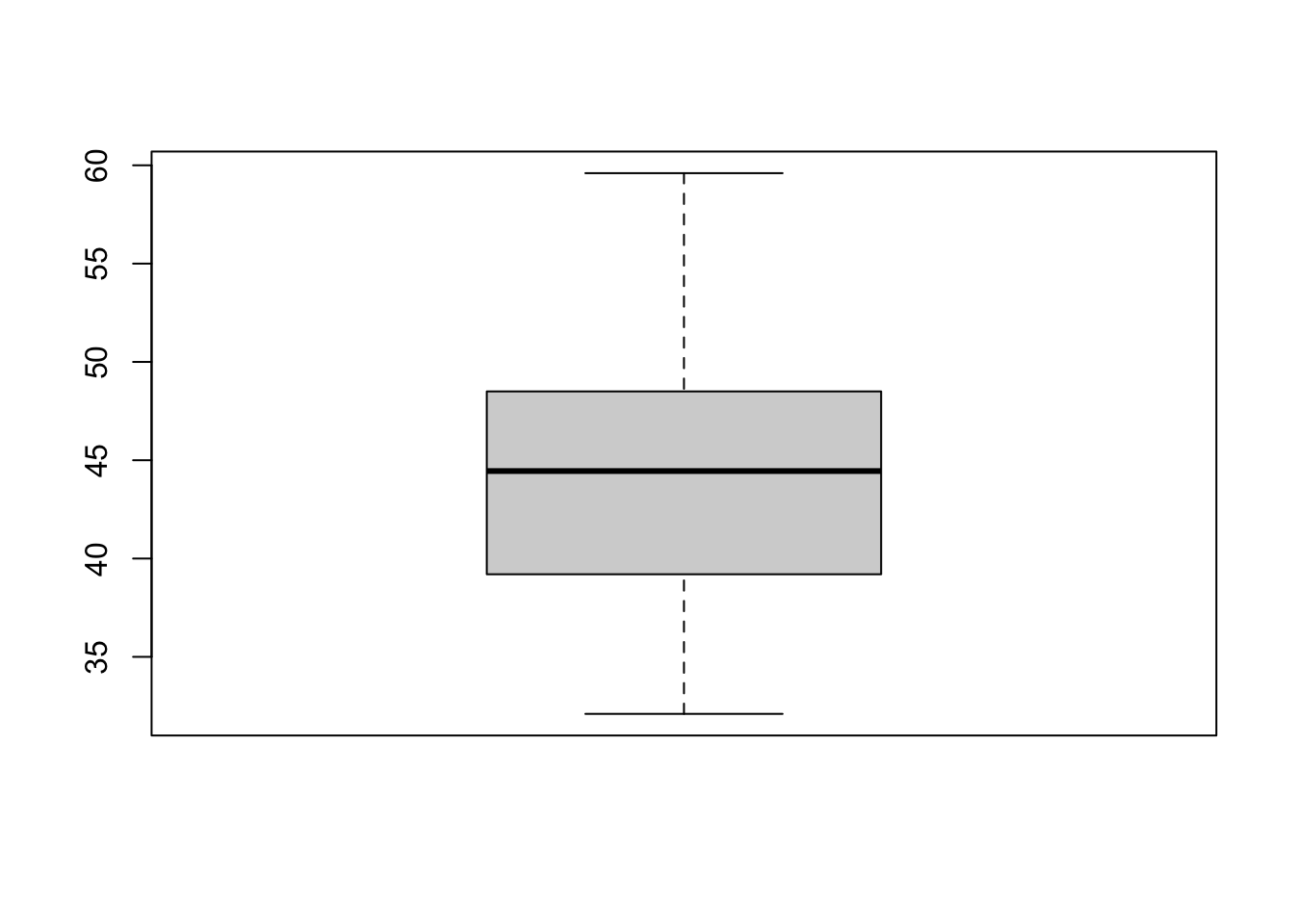
In a boxplot we first have to identify two things: the central box and the external lines (the whiskers). The value indicated by the beginning of the box indicates the first quartile, that is, 25% of the data; then the dark line indicates the median (50% of the data); finally, the top edge of the box marks 75% of the data. In turn, the whiskers extend to reach up to 1.5 the interquartile range, which is the difference between the third and first quartiles of the distribution. All values that are higher or lower than that range will be considered outliers.
If you haven’t fallen asleep with that explanation yet (or the other way around, if you found it difficult to understand), what I want you to take away is this: a boxplot shows you how spread out the sample is. If the box looks compacted and the whiskers are short, it means that most of the data is condensed into close values. In contrast, a boxplot with large boxes and whiskers shows a distribution with greater variation.
We can separate the boxplots with our categorical variables:
boxplot(penguins$bill_length_mm ~ penguins$sex)
To mark groups we use the ~ symbol, very common when defining formulas. In the vast majority of functions in R, you can read the formula as A as a function of B, so in this case it would be the length of the peaks as a function of their sex. The graph shows that: the dispersion of beak lengths depending on whether the penguin is male or female. What we see is that there is sexual dimorphism: females tend to have a shorter beak than males. We can see even more variety if we divide this variable based on the species:
boxplot(penguins$bill_length_mm ~ penguins$species)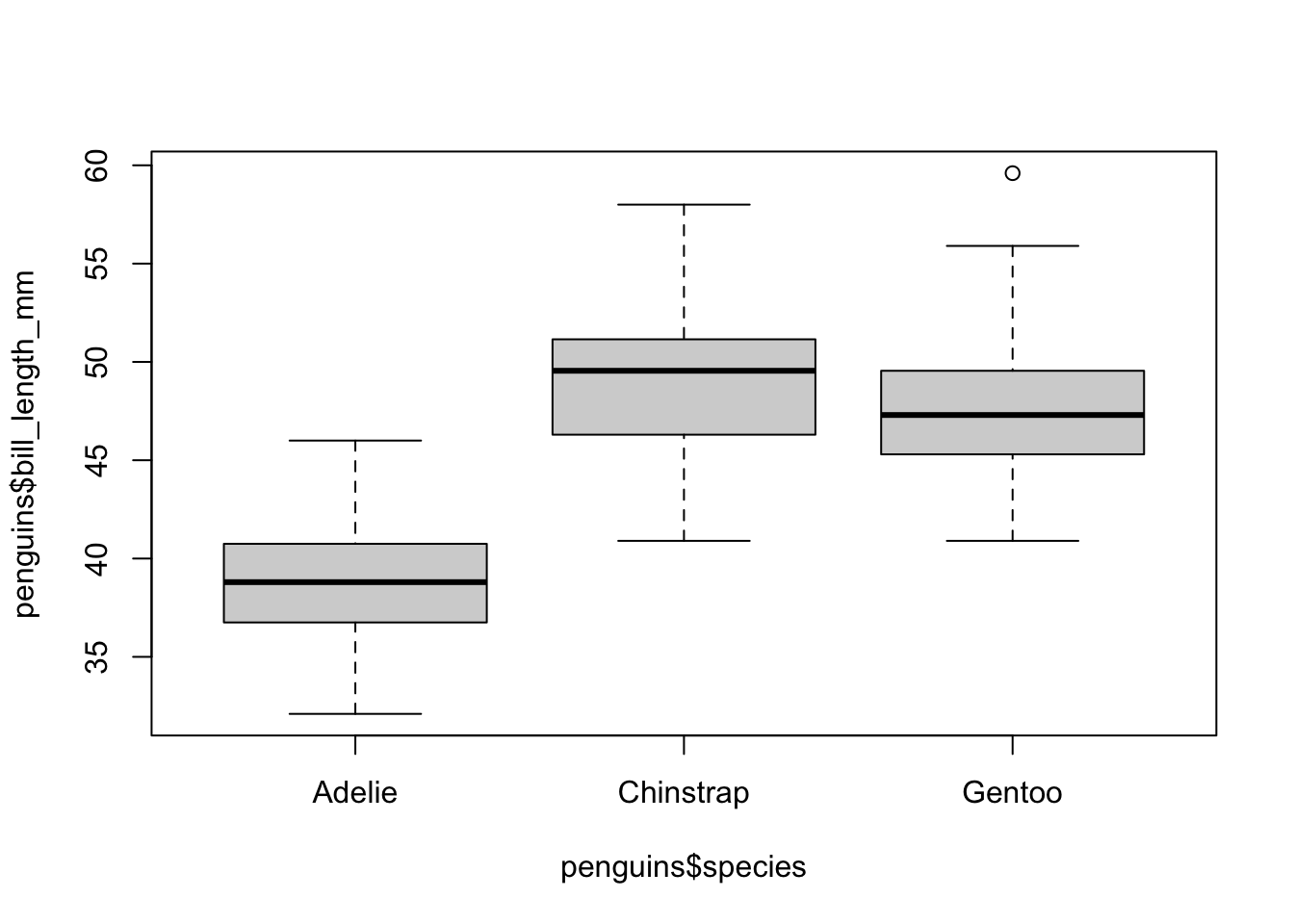
Here we see that the Chinstrap and Gentoo species tend to have beaks of similar lengths (the former a little longer than the latter), while the Adelia penguins have significantly shorter beaks than the others. We can also see a point in the case of Adelie penguins: that point represents a case (a penguin) whose beak length exceeds 1.5 times the interquartile range (i.e., is quite far from the median); this is considered an outlier.
The boxplot is a graph that pairs well with the histogram; Although they transmit similar information, the boxplot is a little more complex and can serve to clarify some intuitions that we may have from the histogram2.
Closing
In this post we saw three classic graphs in the world of research: the histogram, the bar graph and the boxplot. With these graphs you will be able to understand a little more how your data works. As always, remember that you can subscribe to my blog so you don’t miss any updates , and if you have any questions, do not hesitate to contact me. And, if you like what I do, you can buy me a cafecito from Argentina or a kofi.